Question & Answers
The number of turns in secondary coil is 100 times more than that of primary coil. Calculate the output voltage, if 220V is input.
The number of turns in secondary coil is 100 times more than that of primary coil. Calculate the output voltage, if 220V is input. Where is the transformer used ?
In the diagram, C2 = 1/5 of coil of C1. If input voltage is 1200 V, find output V. Give reason for laminating the core .
In the diagram, C2 = 1/5 of coil of C1. If input voltage is 1200 V, find output V. Give reason for laminating the core .
[caption id="attachment_13805" align="alignnone" width="300"]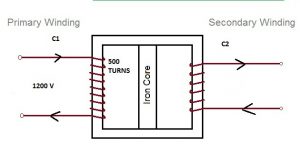 given diagram[/caption]
given diagram[/caption]
The input and output voltage of a transformer are 11000 V and 220 V, find the input current if output is 1650A in transformer.
The input and output voltage of a transformer are 11000 V and 220 V, find the input current if output is 1650A in transformer.
In a transformer, the turn in primary coil is 600 and its emf is 220 V. What will be the number of turns in secondary winding, 60V.
In a transformer, the turn in primary coil is 600 and its emf is 220 V. What will be the number of turns in secondary winding, 60V.
How many turns should be made to produce 110 V of secondary voltage. If primary turns is 3600 and voltage is 440 V.
How many turns should be made to produce 110 V of secondary voltage. If primary turns is 3600 and voltage is 440 V.
A transformer has 220 V primary voltage, 2000 turns. Calculate the secondary turns if voltage is 45 V.
A transformer has 220 V primary voltage, 2000 turns. Calculate the secondary turns if voltage is 45 V.
What is the secondary coil turns while 11 V radio runs in current using 220 V and primary coil 1000 turns.
What is the secondary coil while 11 V radio runs in current using 220 V and primary coil 1000 turns.
Calculate the number of turns in secondary coil, if voltage is 33 V. Primary coil has 200 turns and 220 V.
Calculate the number of turns in secondary coil, if voltage is 33 V. Primary coil has 200 turns and 220 V.
Calculate the secondary coil turn if voltage is 55. Primary voltage is 22 and turns is 1000.
Calculate the secondary coil turn if voltage is 55. Primary voltage is 22 and turns is 1000.
Calculate turns on secondary coil if step up transformer has 220 voltage, 100 turns primary coil. Secondary voltage is 2200 V.
Calculate turns on secondary coil if step up transformer has 220 voltage, 100 turns primary coil. Secondary voltage is 2200 V.
Popular Tags: 7 Days
Upcoming MCQs

Computer Fundamental Multiple Choice Questions Exam Free
- 2022-03-20 12:45
- 60 Mins
- 12 Enrolled
- 25 Full Marks
- 10 Pass Makrs
- 25 Questions